Introduction
Advancing bold, early-stage research by fostering a vibrant, collaborative, cross-disciplinary scientific community.
About
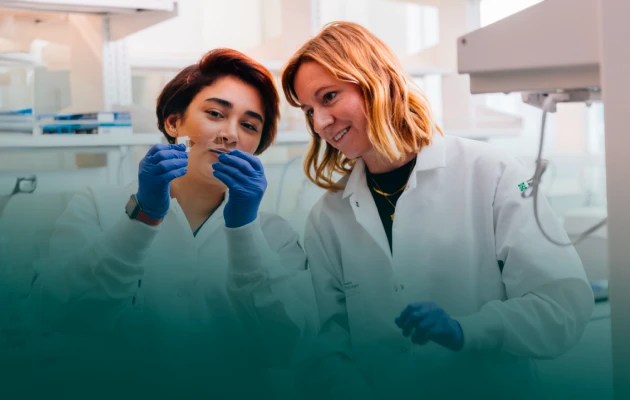
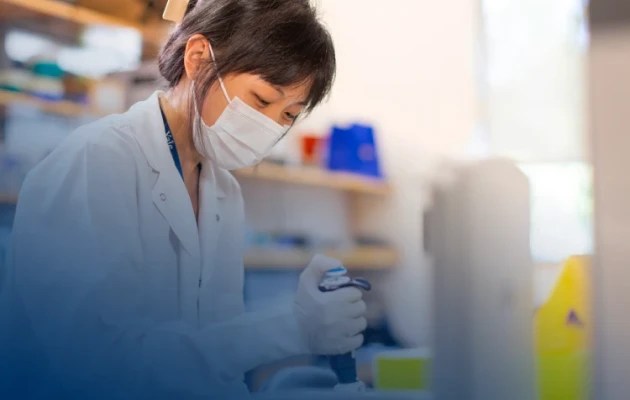
The Investigator Program at Chan Zuckerberg Biohub San Francisco is propelling groundbreaking research by exceptionally creative scientists, engineers, and technologists from Stanford University, UC Berkeley, and UC San Francisco. Over a five-year term, each Investigator receives $1 million of unrestricted funding, giving them the freedom to pursue their boldest, most visionary ideas. These daring projects accelerate the pace of scientific discovery and help CZ Biohub realize its vision of curing, preventing, or managing every disease by the end of the century.
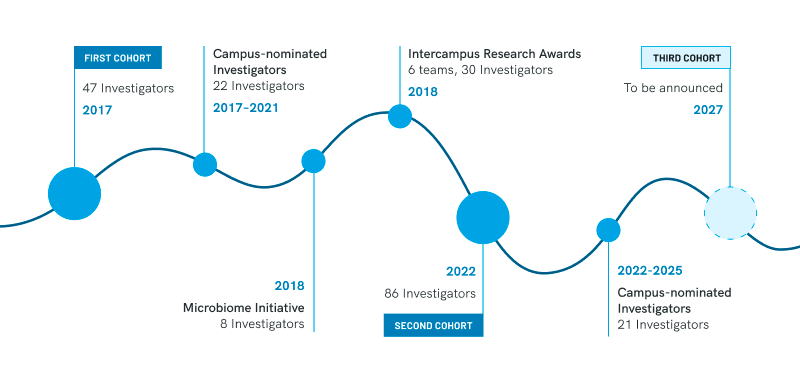
Awardees were selected after a competitive process and represent a diverse array of disciplines. Additional Investigators were named following a nomination by their university in a program that provided resources for recruiting talented new faculty and boosting the work of early- to mid-career faculty.

In addition to receiving funding in support of their research, Investigators gather in person every two weeks to learn from, inspire, and find opportunities to collaborate with one another. The program is designed to catalyze connections and cultivate a rich culture of scientific exchange.
And twice a year, the SF Investigator Program holds an Inter-lab Confab, with keynotes, lightning talks, poster sessions, and time for networking. These events foster opportunities for the wider CZ Biohub SF community, including — and featuring — students, postdocs, and staff scientists from Biohub and the labs of Biohub Investigators at our partner institutions, to talk science, learn about one another’s work, help share expertise, solve challenges, and establish fresh collaborations.
Our Impact
Explore stories about Biohub San Francisco Investigators.
Investigator Voices
Program Staff

Bill Burkholder
Director, Scientific Program Management

Ben Warner
Senior Program and Scientific Event Manager

Catherine Tan
Program Impact Analyst